MASON (Make AntiSense Oligomers Now) is a web-tool which guides the design of bacterial antisense oligonucleotides (ASOs). We initially developed it for peptide nucleic acids (PNAs), although it can also be used for designing other ASOs such as PMOs or LNAs. All you need is a genome and the gene you want to target.
Read our paper: RNA Journal - Design and off-target prediction for antisense oligomers targeting bacterial mRNAs with the MASON webserver public
Report bugs/problems by opening a GitHub issue: github.com/BarquistLab/mason/issuesbug_report
Contact us via email: jakobjung@tutanota.com email
Start designing ASOs for targeting any bacterial gene: start MASON fast_forward
MASON update - MASON 2.0
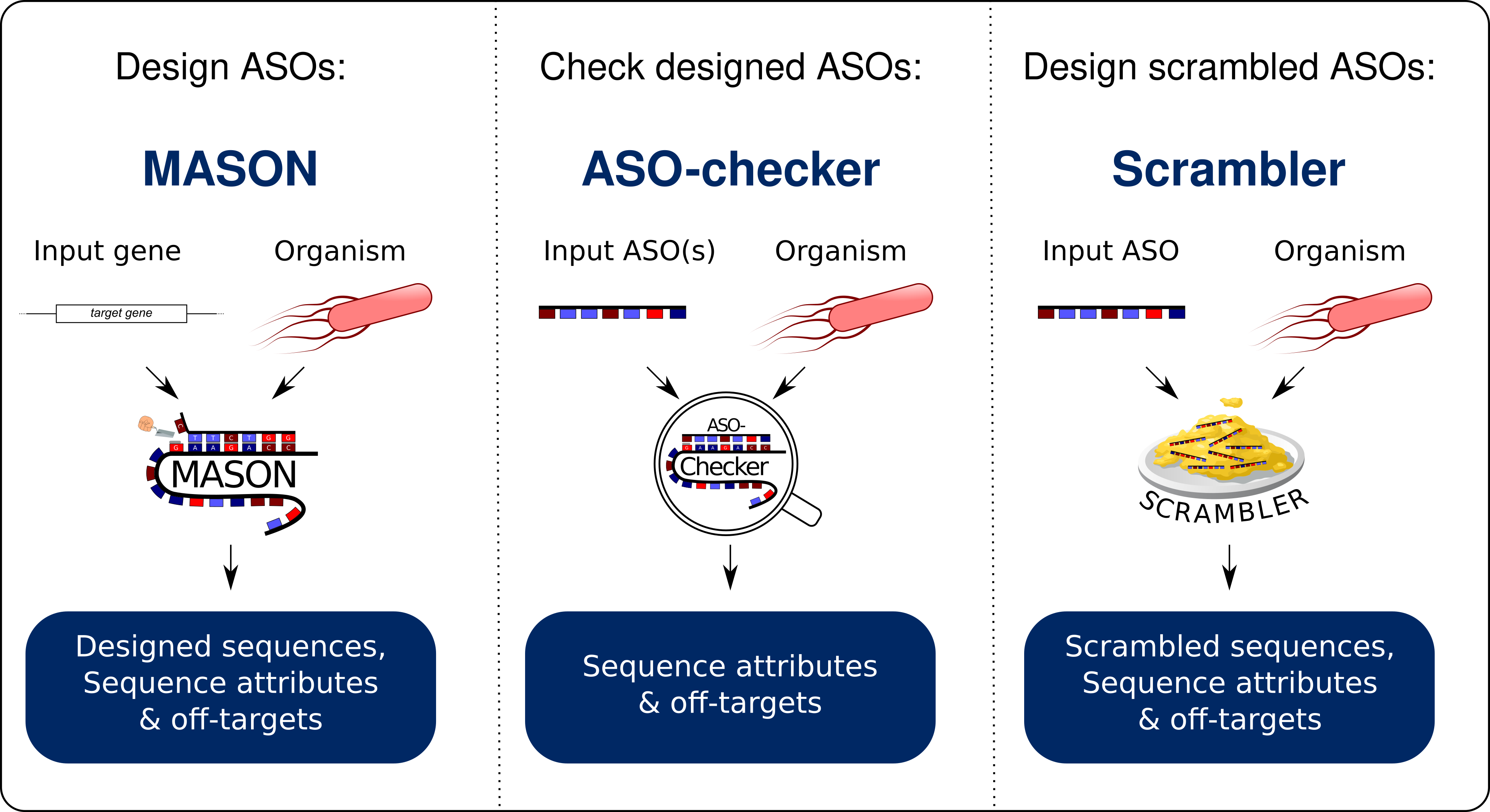
We have updated the MASON webserver with two new features: ASO-checker and Scrambler. ASO-checker validates your ASO design by checking for off-targets in the genome while checking for different ASO-specific features. Scrambler scrambles the sequence of your ASO to generate the best negative control ASO sequence for your experiment. Additionally, MASON 2.0 introduces:
- NCBI accession download:enter an NCBI assembly accession to automatically download genome and annotation files, no manual upload needed
- 10 preset organisms with pre-loaded genomes and essential gene sets
- Essential gene off-target screening:screen ASO off-targets against essential genes
- RNA secondary structure visualization:VARNA plots of the target region with Shine-Dalgarno highlighting
- Improved Shine-Dalgarno detection:wider search window with extended SD motif recognition
- Self-binding energy (ΔG):MFE-based self-complementarity assessment for each ASO
- Off-target breakdown by mismatch level:off-targets are now reported separately for 0, 1, 2, and 3 mismatches
- ML-based MIC prediction BETA:optional random forest model to rank ASO efficacy
There are now three tools available on the MASON homepage: